Easy to use systems biology suite for your Mac #Numerical algorithm #SBML simulation #Run simulation #Algorithm #Systems biology #SBML
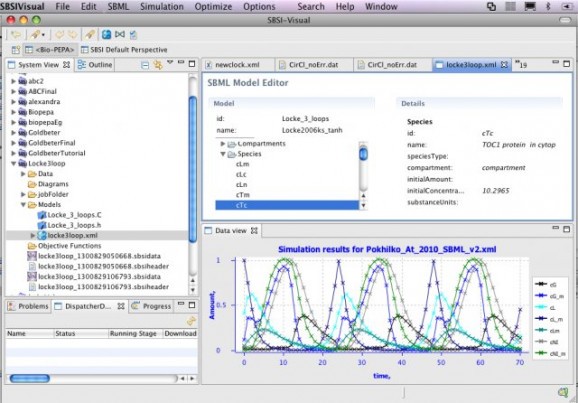
SBSI (Systems Biology Software Infrastructure) is an open source, easy to use suite of tools for systems biology, such as parallelized numerical algorithms, and an Eclipse RCP based client for visualizing results, running simulations, and integrating SBML based plugins.
There are three main software components in SBSI: SBSINumerics, SBSIVisual and SBSIDispatcher.
SBSINumerics contains a configurable, parallelized command line application for running parameter estimations.
SBSIVisual is a desktop application, that provides easier access to SBSINumerics functionality for non-programmers, and can display results, run simulations, and interact with systems biology databases.
SBSIDispatcher is a middleware component, that can deploy and manage SBSINumerics procedures on backend servers.
What's new in SBSI 1.4.5:
- Minor update with bug-fixes
SBSI 1.4.5
add to watchlist add to download basket send us an update REPORT- runs on:
- Mac OS X 10.5 or later (PPC & Intel)
- file size:
- 123.7 MB
- main category:
- Math/Scientific
- developer:
- visit homepage
ShareX
4k Video Downloader
IrfanView
7-Zip
calibre
Windows Sandbox Launcher
Zoom Client
Context Menu Manager
Bitdefender Antivirus Free
Microsoft Teams
- Context Menu Manager
- Bitdefender Antivirus Free
- Microsoft Teams
- ShareX
- 4k Video Downloader
- IrfanView
- 7-Zip
- calibre
- Windows Sandbox Launcher
- Zoom Client