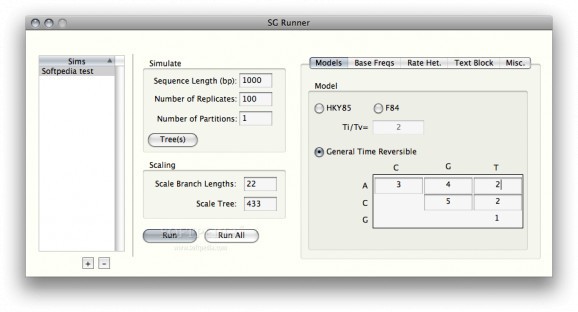
Simulates the evolution of nucleotide or amino acid sequences along a phylogeny. #Nucleotide evolution #Evolution simulator #Simulate evolution #Simulate #Evolution #Simulator
Seq-Gen is an application that will simulate the evolution of nucleotide or amino acid sequences along a phylogeny, using common models of the substitution process.
Also, a range of models of molecular evolution will be implemented including the general reversible model.
State frequencies and other parameters of the model may be given and site-specific rate heterogeneity may also be incorporated in a number of ways. Any number of trees may be read in and the program will produce any number of data sets for each tree.
Thus large sets of replicate simulations can be easily created. It has been designed to be a general purpose simulator that incorporates most of the commonly used (and computationally tractable) models of molecular sequence evolution.
What's new in Seq-Gen 1.3.3:
- New features:
- MtArt amino acid model added by Lars Jermiin.
- Bugs fixed:
Seq-Gen 1.3.3
add to watchlist add to download basket send us an update REPORT- runs on:
- Mac OS X (PPC & Intel)
- file size:
- 61 KB
- main category:
- Math/Scientific
- developer:
- visit homepage
calibre
Bitdefender Antivirus Free
4k Video Downloader
Context Menu Manager
Windows Sandbox Launcher
Microsoft Teams
IrfanView
ShareX
Zoom Client
7-Zip
- ShareX
- Zoom Client
- 7-Zip
- calibre
- Bitdefender Antivirus Free
- 4k Video Downloader
- Context Menu Manager
- Windows Sandbox Launcher
- Microsoft Teams
- IrfanView