Cross-platform tool for biological rule-based modeling, simulation, and visualization. #Biological modeller #Biochemistry simulation #Biology visualizer #Biology #Modeller #Simulator
First of all, modern biological experiments generate great amounts of data describing intracellular dynamics.
The rule-based languages for describing intracellular biochemistry allow for the construction and simulation of detailed models with unprecedented scope and precision.
We can use rule-based Models to suggest new hypotheses and new ideas for future experimentation. Rule-based models can be difficult to understand and time consuming to debug.
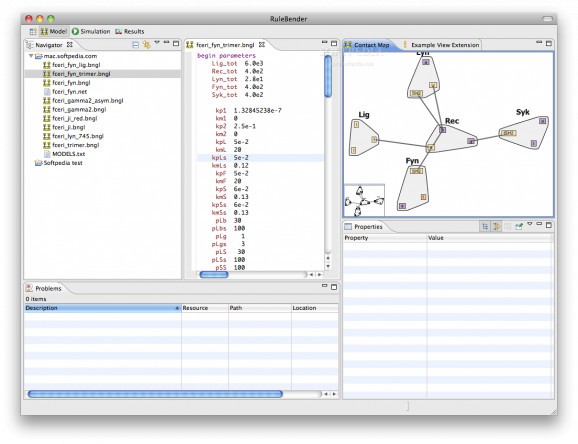
The RuleBender application is a free utility that allows you to construct, debug, simulate and analyze rule-based biological models in the BioNetGen language.
Detailed instructions on how to use the RuleBender utility on your Mac are available HERE.
RuleBender is cross-platform and it works on Mac OS X, Windows and Linux. Binaries for the Windows and Linux platforms are available on the project's homepage.
System requirements
RuleBender 2.0.382
add to watchlist add to download basket send us an update REPORT- runs on:
- Mac OS X 10.5 or later (Intel only)
- file size:
- 100.6 MB
- filename:
- RuleBender-2.0.382-osx64.zip
- main category:
- Math/Scientific
- developer:
- visit homepage
ShareX
Windows Sandbox Launcher
Bitdefender Antivirus Free
calibre
Zoom Client
Context Menu Manager
Microsoft Teams
4k Video Downloader
7-Zip
IrfanView
- 4k Video Downloader
- 7-Zip
- IrfanView
- ShareX
- Windows Sandbox Launcher
- Bitdefender Antivirus Free
- calibre
- Zoom Client
- Context Menu Manager
- Microsoft Teams