Proteomic data analysis tool in Java. #Proteomic data analysis #Analyze Proteomic data #Peptide identifier #Proteomics #Peptide #Analyzer
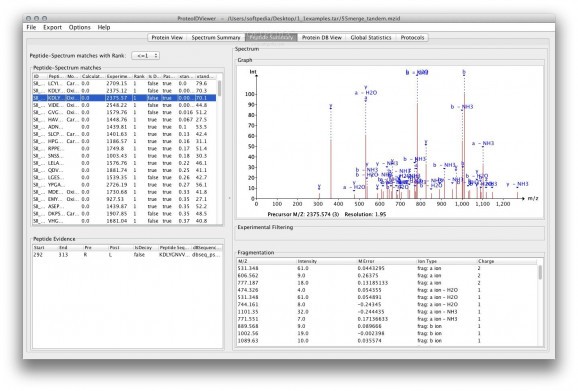
ProteoIDViewer is a free and open-source GUI that allows the experimental proteomics community to use the mzIdentML standard.
ProteoIDViewer aims to simplify proteomics data sharing for laboratory scientists to solve one of the main difficulties to open proteomic data analysis and large scale data sharing (http://code.google.com/p/mzidentml-viewer/).
The laboratory scientists are the main users of the mzIdentML Viewer which does not require bioinformatics support to get started. The viewer can be simply downloaded and installed with no complex setup procedures.
ProteoIDViewer provides intuitive and useful views of peptides/proteins and the search metadata. moreover, the software calls any of the functions within mzIdentML Library, and exporters to spreadsheets.
ProteoIDViewer is a cross-platform utility capable of running on any operating system that comes with Java support (e.g. Mac OS X, Windows, Linux).
System requirements
ProteoIDViewer 1.2
add to watchlist add to download basket send us an update REPORT- runs on:
- Mac OS X (-)
- file size:
- 23.8 MB
- filename:
- ProteoIDViewer-1.2.zip
- main category:
- Math/Scientific
- developer:
- visit homepage
4k Video Downloader
paint.net
Windows Sandbox Launcher
IrfanView
7-Zip
ShareX
Microsoft Teams
Bitdefender Antivirus Free
Zoom Client
calibre
- Bitdefender Antivirus Free
- Zoom Client
- calibre
- 4k Video Downloader
- paint.net
- Windows Sandbox Launcher
- IrfanView
- 7-Zip
- ShareX
- Microsoft Teams