Tool to explore the topology of membrane proteins. #Protein topology #Membrane protein #Study protein #Protein #Study #Topology
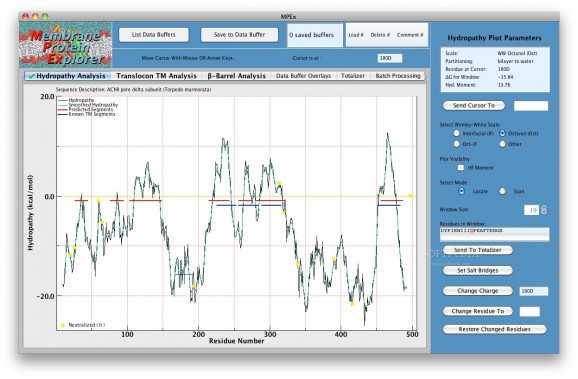
MPEx is a free and easy to use tool for exploring the topology and other features of membrane proteins by means of hydropathy plots based upon thermodynamic and biological principles.
MPEx is designed for examination of membrane protein sequences using hydropathy-plot methods popularized by Kyte and Doolittle (1982).
That is, it is based on sliding-window analysis that represents an amino acid sequence as a sequence of numbers representing physical or statistical properties of the amino acid sequence.
The most common physical property is hydropathy, which is based upon the free energy of partitioning of amino acids between water and membrane.
This version of MPEx uses two types of hydropathy scales: Experiment-based whole-residue partitioning scales determined in this laboratory and experiment-based biological partitioning scales determined in collaboration with the von Heijne laboratory at Stockholm University.
System requirements
MPEx 3.2
add to watchlist add to download basket send us an update REPORT- runs on:
- Mac OS X (-)
- file size:
- 1 KB
- main category:
- Math/Scientific
- developer:
- visit homepage
7-Zip
Microsoft Teams
Windows Sandbox Launcher
Zoom Client
calibre
Context Menu Manager
Bitdefender Antivirus Free
4k Video Downloader
IrfanView
ShareX
- 4k Video Downloader
- IrfanView
- ShareX
- 7-Zip
- Microsoft Teams
- Windows Sandbox Launcher
- Zoom Client
- calibre
- Context Menu Manager
- Bitdefender Antivirus Free