Open source and highly flexible versatile cross-platform utility designed from the ground up in order to analyze and qualify biological images #Image analysis #Image quantifier #Biology analysis #Biology #Analysis #Phenotype
Moreover, CellProfiler is designed for modular, flexible, high-throughput analysis of images, measuring a wide variety of properties from size, shape, and intensity to the texture of every cell (or other object) in every image.
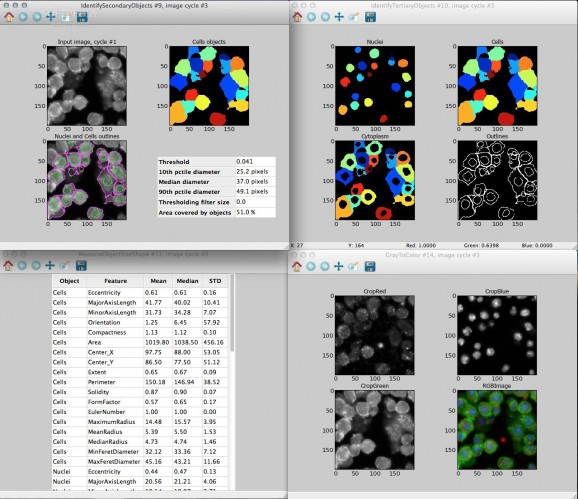
Using the point-and-click graphical user interface (GUI), users construct an image analysis "pipeline", a sequential series of modules that each perform an image processing function such as illumination correction, object identification (segmentation), and object measurement.
Users mix and match modules and adjust their settings to measure the phenotype of interest.
While originally designed for high-throughput images, it is equally appropriate for low-throughput assays as well (i.e., assays of < 100 images).
What's new in CellProfiler 4.2.6:
- There are no changes to the functional CellProfiler codebase in this release compared to 4.2.5; the only changes are changes intended to make it easier to pip install CellProfiler by updating some pins that did not have wheels on some modern systems, and pinning at the top of their range packages whose updates break existing functionality. If you are using CellProfiler downloaded from our website, there is no need to update; these changes should only affect those installing via a local Python environment.
- PRs:
- Tighten some pins, loosen others by @bethac07 in #4801
CellProfiler 4.2.6
add to watchlist add to download basket send us an update REPORT- runs on:
- macOS 10.6.8 or later (Intel only)
- file size:
- 491.3 MB
- filename:
- CellProfiler-macOS-4.2.6.zip
6 screenshots:
- main category:
- Math/Scientific
- developer:
- visit homepage
Microsoft Teams
Effortlessly chat, collaborate on projects, and transfer files within a business-like environment by employing this Microsoft-vetted application
Bitdefender Antivirus Free
Feather-light and free antivirus solution from renowned developer that keeps the PC protected at all times from malware without requiring user configuration
ShareX
Capture your screen, create GIFs, and record videos through this versatile solution that includes various other amenities: an OCR scanner, image uploader, URL shortener, and much more
calibre
Effortlessly keep your e-book library thoroughly organized with the help of the numerous features offered by this efficient and capable manager
Context Menu Manager
Customize Windows’ original right-click context menu using this free, portable and open-source utility meant to enhance your workflow
4k Video Downloader
Export your favorite YouTube videos and playlists with this intuitive, lightweight program, built to facilitate downloading clips from the popular website
IrfanView
With support for a long list of plugins, this minimalistic utility helps you view images, as well as edit and convert them using a built-in batch mode
7-Zip
An intuitive application with a very good compression ratio that can help you not only create and extract archives, but also test them for errors
Zoom Client
The official desktop client for Zoom, the popular video conferencing and collaboration tool used by millions of people worldwide
Windows Sandbox Launcher
Set up the Windows Sandbox parameters to your specific requirements, with this dedicated launcher that features advanced parametrization
% discount
7-Zip
- 7-Zip
- Zoom Client
- Windows Sandbox Launcher
- Microsoft Teams
- Bitdefender Antivirus Free
- ShareX
- calibre
- Context Menu Manager
- 4k Video Downloader
- IrfanView
essentials
Click to load comments
This enables Disqus, Inc. to process some of your data. Disqus privacy policy